Generic optimizer that can be customized by user provided functions for generating shuffles and progressing towards the minimal score
Source:R/optimize.R
optimize_design.RdGeneric optimizer that can be customized by user provided functions for generating shuffles and progressing towards the minimal score
optimize_design(
batch_container,
samples = NULL,
scoring = NULL,
n_shuffle = NULL,
shuffle_proposal_func = NULL,
acceptance_func = accept_strict_improvement,
aggregate_scores_func = identity,
check_score_variance = TRUE,
autoscale_scores = FALSE,
autoscaling_permutations = 100,
autoscale_useboxcox = TRUE,
sample_attributes_fixed = FALSE,
max_iter = 10000,
min_delta = NA,
quiet = FALSE
)Arguments
- batch_container
An instance of
BatchContainer.- samples
A
data.framewith sample information. Should beNULLif theBatchContaineralready has samples in it.- scoring
Scoring function or a named
list()of scoring functions.- n_shuffle
Vector of length 1 or larger, defining how many random sample swaps should be performed in each iteration. If
length(n_shuffle)==1, this sets no limit to the number of iterations. Otherwise, the optimization stops if the swapping protocol is exhausted.- shuffle_proposal_func
A user defined function to propose the next shuffling of samples. Takes priority over n_shuffle if both are provided. The function is called with a BatchContainer
bcand an integer parameteriterationfor the current iteration number, allowing very flexible shuffling strategies. Mapper syntax is supported (seepurrr::as_mapper()). The returned function must either return a list with fieldssrcanddst(for pairwise sample swapping) or a numeric vector with a complete re-assigned sample order.- acceptance_func
Alternative function to select a new score as the best one. Defaults to strict improvement rule, i.e. all elements of a score have to be smaller or equal in order to accept the solution as better. This may be replaced with an alternative acceptance function included in the package (e.g.
mk_simanneal_acceptance_func()) or a user provided function. Mapper syntax is supported (seepurrr::as_mapper()).- aggregate_scores_func
A function to aggregate multiple scores AFTER (potential) auto-scaling and BEFORE acceptance evaluation. If a function is passed, (multi-dimensional) scores will be transformed (often to a single double value) before calling the acceptance function. E.g., see
first_score_only()orworst_score(). Note that particular acceptance functions may require aggregation of a score to a single scalar in order to work, see for example those generated bymk_simanneal_acceptance_func(). Mapper syntax is supported (seepurrr::as_mapper()).- check_score_variance
Logical: if TRUE, scores will be checked for variability under sample permutation and the optimization is not performed if at least one subscore appears to have a zero variance.
- autoscale_scores
Logical: if TRUE, perform a transformation on the fly to equally scale scores to a standard normal. This makes scores more directly comparable and easier to aggregate.
- autoscaling_permutations
How many random sample permutations should be done to estimate autoscaling parameters. (Note: minimum will be 20, regardless of the specified value)
- autoscale_useboxcox
Logical; if TRUE, use a boxcox transformation for the autoscaling if possible at all. Requires installation of the
bestNormalizepackage.- sample_attributes_fixed
Logical; if TRUE, sample shuffle function may generate altered sample attributes at each iteration. This affects estimation of score distributions. (Parameter only relevant if shuffle function does introduce attributes!)
- max_iter
Stop optimization after a maximum number of iterations, independent from other stopping criteria (user defined shuffle proposal or min_delta).
- min_delta
If not NA, optimization is stopped as soon as successive improvement (i.e. euclidean distance between score vectors from current best and previously best solution) drops below min_delta.
- quiet
If TRUE, suppress non-critical warnings or messages.
Value
A trace object
Examples
data("invivo_study_samples")
bc <- BatchContainer$new(
dimensions = c("plate" = 2, "column" = 5, "row" = 6)
)
bc <- optimize_design(bc, invivo_study_samples,
scoring = osat_score_generator("plate", "Sex"),
max_iter = 100
)
#> Warning: NAs in features / batch columns; they will be excluded from scoring
#> Checking variances of 1-dim. score vector.
#> ... (141.154) - OK
#> Initial score: 182.5
#> Achieved score: 132.5 at iteration 1
#> Achieved score: 90.5 at iteration 2
#> Achieved score: 56.5 at iteration 6
#> Achieved score: 30.5 at iteration 11
#> Achieved score: 12.5 at iteration 15
#> Achieved score: 2.5 at iteration 16
#> Achieved score: 0.5 at iteration 31
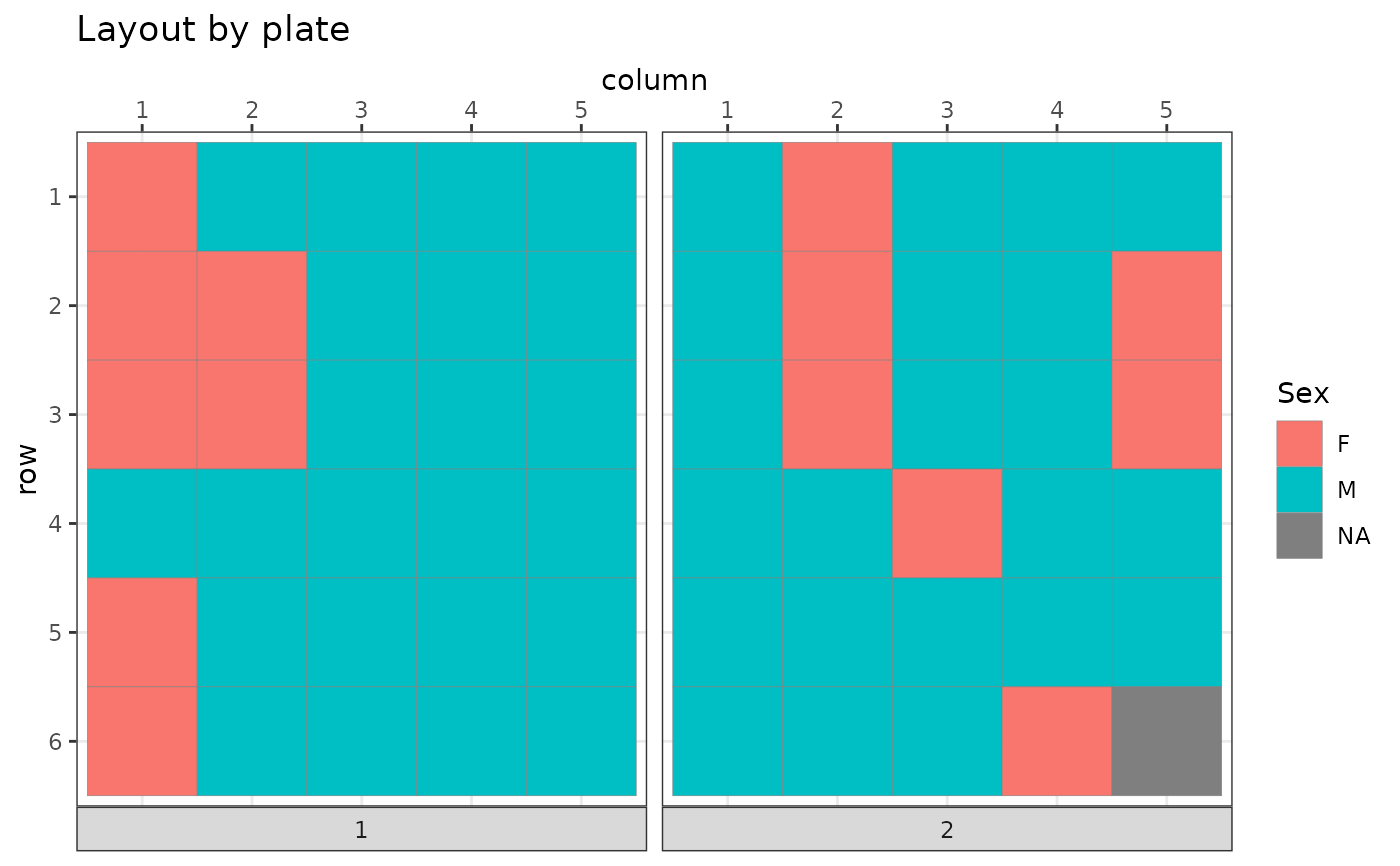
plot_plate(bc$get_samples(), .col = Sex)